t tests — make sure you know every detail of these
- including
When true: population variances are equal. Called Pooled -Test. Use on fail 2 rej null hypothesis
t.test(group1_data, group2_data,
var.equal = TRUE,
alternative = "two.sided")When false Welch -Test. safer choice unless there is strong evidence for equal variances Use when reject null hypothesis from var.test. Same as above, but false.
NULL and Alternate hypotheses
- : Represents a statement of “no effect,” “no difference,” or “no relationship”
- always includes an equality sign (, , or ).
- what you’re trying to find.
t.test(Group\_A, Group\_B, alternative = "\text{less}")Using data in R to create tests
Predictions
SLR:
- expected value of the response variable for a specific value p of the predictor variable - mean response of the population when the predictor variable is fixed at -
- Least Squares estimates
- Find values for the slope () and the intercept () that minimize Sum of Squared Errors (SSE) use
a = lm(y_data ~ x_data)
summary(a)- ci confidence interval creation in r
# Find the 95% Confidence Intervals for the Intercept and Slope
confint(my_model, level = 0.95)Use s20x package
MGF
Packages all raw moments of random variable into an expr. 2 ways to find (vaience) and (mean). To find -th raw moment, take derative a@ 0
- Find : calculated using the second and first raw moments:
- Identify distributions
Gaps
(or stuff)
- (or )
- (or )
Power
Probability of correctly rejecting when true aka
power_result <- power.t.test(
n = 30, # Sample size
delta = 1, # The true difference in means (Effect Size)
sd = 2, # Population standard deviation
sig.level = 0.05, # Significance level (alpha)
type = "one.sample",# Type of t-test
alternative = "two.sided" # Hypothesis direction
)Calculate probabilities in R
LSE and varience
make predictions using the estimated regression line
- LSE for slope
- Sample estimate of the slope, calculated by dividing the sample covariance of and (scaled by ) by the sample variance of (scaled by ).
- defines the steepness of the estimated regression line
- Varience of LSE for intercept
- calculate the Standard Error of , which is then used in the -test for and the CI for .
my_model <- lm(y_data ~ x_data)
coefficients(my_model)
beta_0_hat <- coefficients(my_model)[1] # The Intercept
beta_1_hat <- coefficients(my_model)[2] # The Slope (x_data)Show the estimators are unbiased
unbiased if its expected value is equal to the true population parameter being estimated:
Find their variances etc
t-tests:
- Can you derive results by hand (P-values, t values, cis)
Perform tests using the three methods: CI, P values, RAR
Setup: We’re already given the output from t.test, and need to confirm the rest of the things Testing for population mean :
- Confidence interval:
- Reject if the null value () is not in CI
To get bounds:
estimate <- 7.9140
std_error <- 0.2471
t_critical <- 2.048
lower_bound <- estimate - t_critical * std_error
upper_bound <- estimate + t_critical * std_error
#which are our bounds..- P-value:
- Fail to Reject if the P-value from t.test
- Rejection/Acceptance Region (RAR):
- Fail to Reject if the calculated test statistic () falls into the acceptance region (i.e., ).
# RAR example
alpha <- 0.05 df <- 28 # From the SLR summary example in the file
t_critical <- qt(1 - alpha/2, df)
#then if t_calc is 3.0, and t_crit = 2, reject H_0
- When use a ?
- conduct two-sample independent -test
Estimation:
- Max Lik: can you perform these calculations and proofs?
- Bootstrap — do you understand the code?
# Basic Bootstrap for a mean in R
n <- length(my_data)
B <- 10000 # Number of resamples
results <- numeric(B)
for(i in 1:B) {
# Resample with replacement
resample <- sample(my_data, size = n, replace = TRUE)
results[i] <- mean(resample)
}
# Find the Bootstrap Confidence Interval
quantile(results, c(0.025, 0.975))- Interval estimation: can you derive results that we did in class?
- Expected Value :
- For Exponential data, , so
- Variance :
- Assuming independence,
- For Exponential data, , so
- Distribution:
- By the Central Limit Theorem, if is large, will be approximately Normal11.
Regression:
#use
a= lm()
summary(a)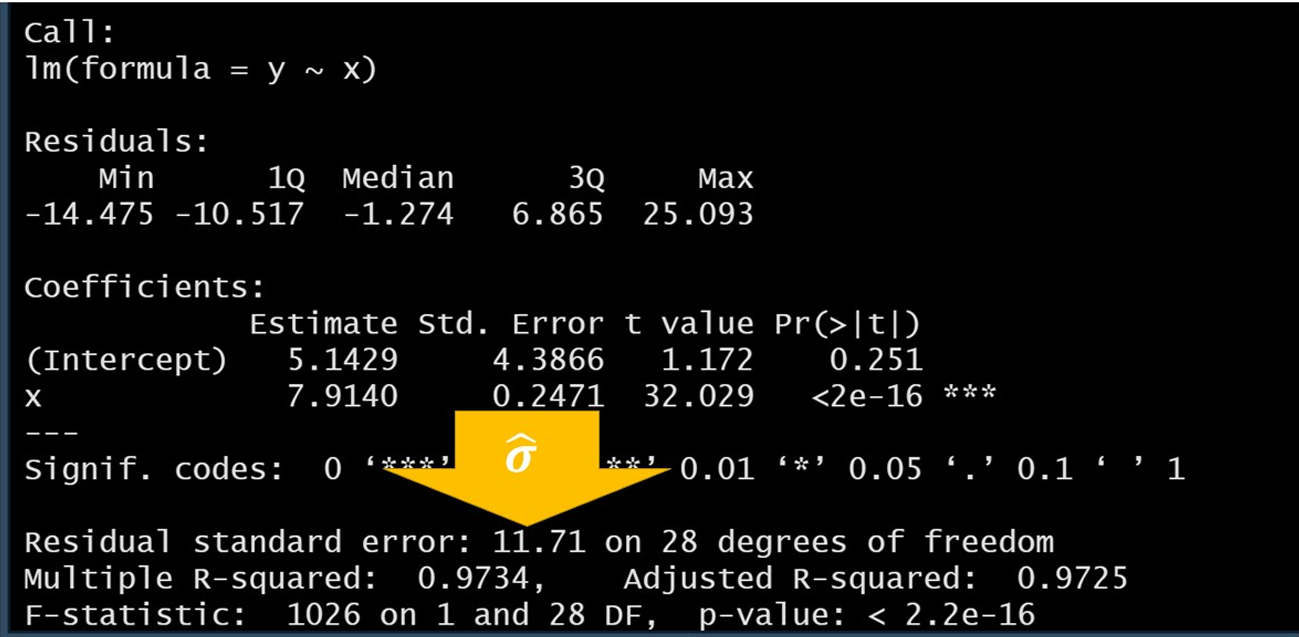
- (Intercept) = 5.14…
- (Slope) = 7.91..
- for SLR sample size = DOF +2 (30 in this case)
- Residual Standard Error (): 11.71
- Multiple R-squared: 97.3% of the variation in is explained by the linear relationship with
- F-statistic: p-value is very small, we reject the null hypothesis
- considered small if it is less than or equal to
- verify t-vales:
- dividing the Estimate by the Std. Error in
xrow.
- dividing the Estimate by the Std. Error in
- Find t_crit:
t_crit <- qt(0.975, 28)# 28 = dfm 0.975 for 95% conf interval